nSTAT Python Paper Examples
This page mirrors the MATLAB paper-example index for the standalone Python port.
Canonical source files:
examples/paper/*.pynstat/paper_examples_full.py
Run Everything
python tools/paper_examples/build_gallery.py
Outputs:
Figure metadata:
docs/figures/manifest.jsonGallery page:
docs/paper_examples.mdFigures:
docs/figures/example01/…docs/figures/example05/
Example Index
ID |
Thumbnail |
Standalone source |
Question |
Run command |
Figure gallery |
|---|---|---|---|---|---|
|
|
Do mEPSCs follow constant vs piecewise Poisson firing under Mg2+ washout? |
|
||
|
|
How do explicit whisker stimulus and spike history improve thalamic GLM fits? |
|
||
|
|
How do PSTH and SSGLM capture within-trial and across-trial dynamics? |
|
||
|
|
Which receptive-field basis (Gaussian vs Zernike) better fits place cells? |
|
||
|
|
How well do adaptive/hybrid point-process filters decode stimulus and reach state? |
|
Gallery
Example 01: mEPSC Poisson Models Under Constant and Washout Magnesium
Question: Do mEPSCs follow constant vs piecewise Poisson firing under Mg2+ washout?
Run command: python examples/paper/example01_mepsc_poisson.py
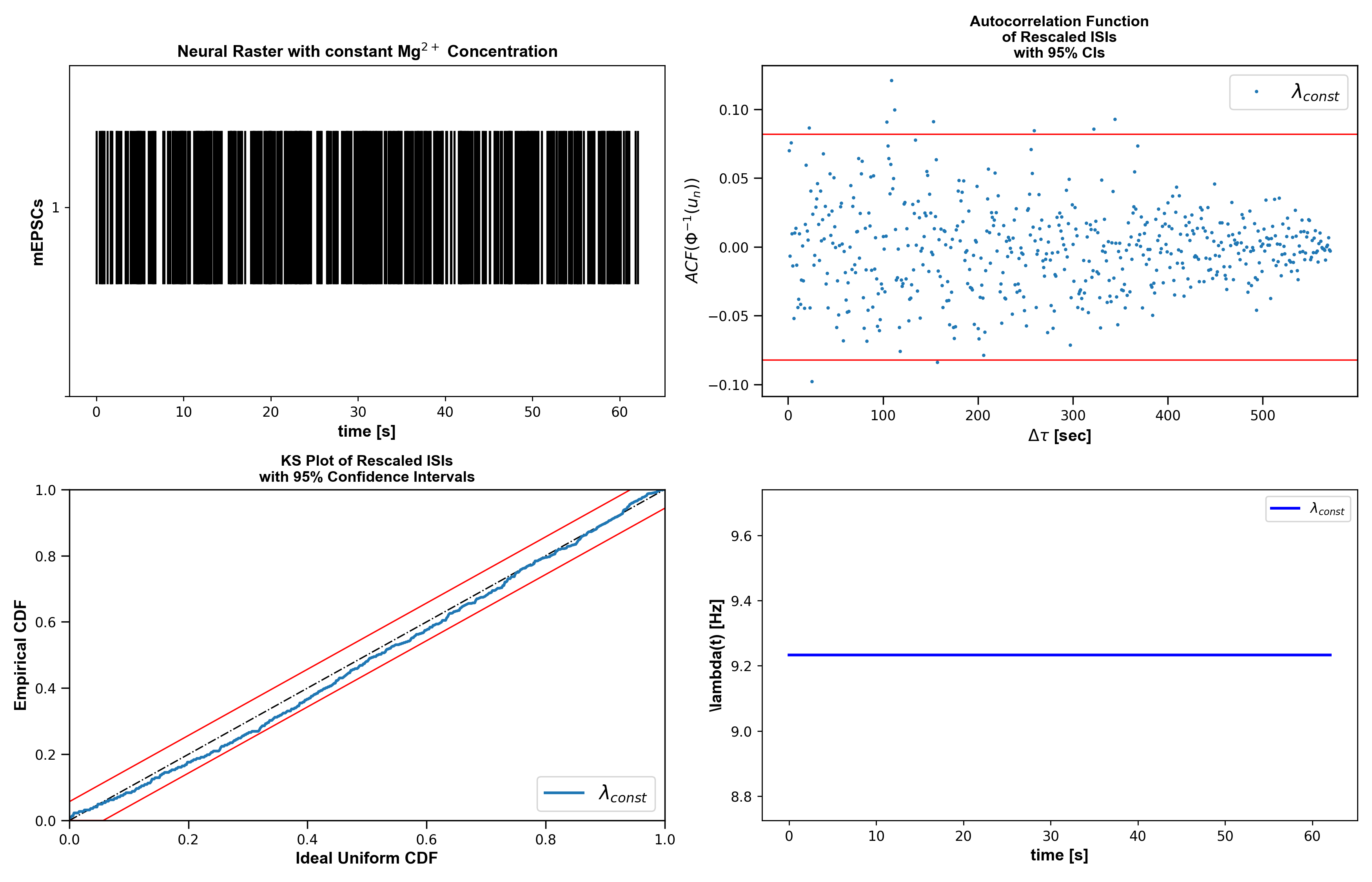
Expected figure files:
docs/figures/example01/fig01_constant_mg_summary.pngdocs/figures/example01/fig02_washout_raster_overview.pngdocs/figures/example01/fig03_piecewise_baseline_comparison.png
Example 02: Whisker Stimulus GLM With Lag and History Selection
Question: How do explicit whisker stimulus and spike history improve thalamic GLM fits?
Run command: python examples/paper/example02_whisker_stimulus_thalamus.py
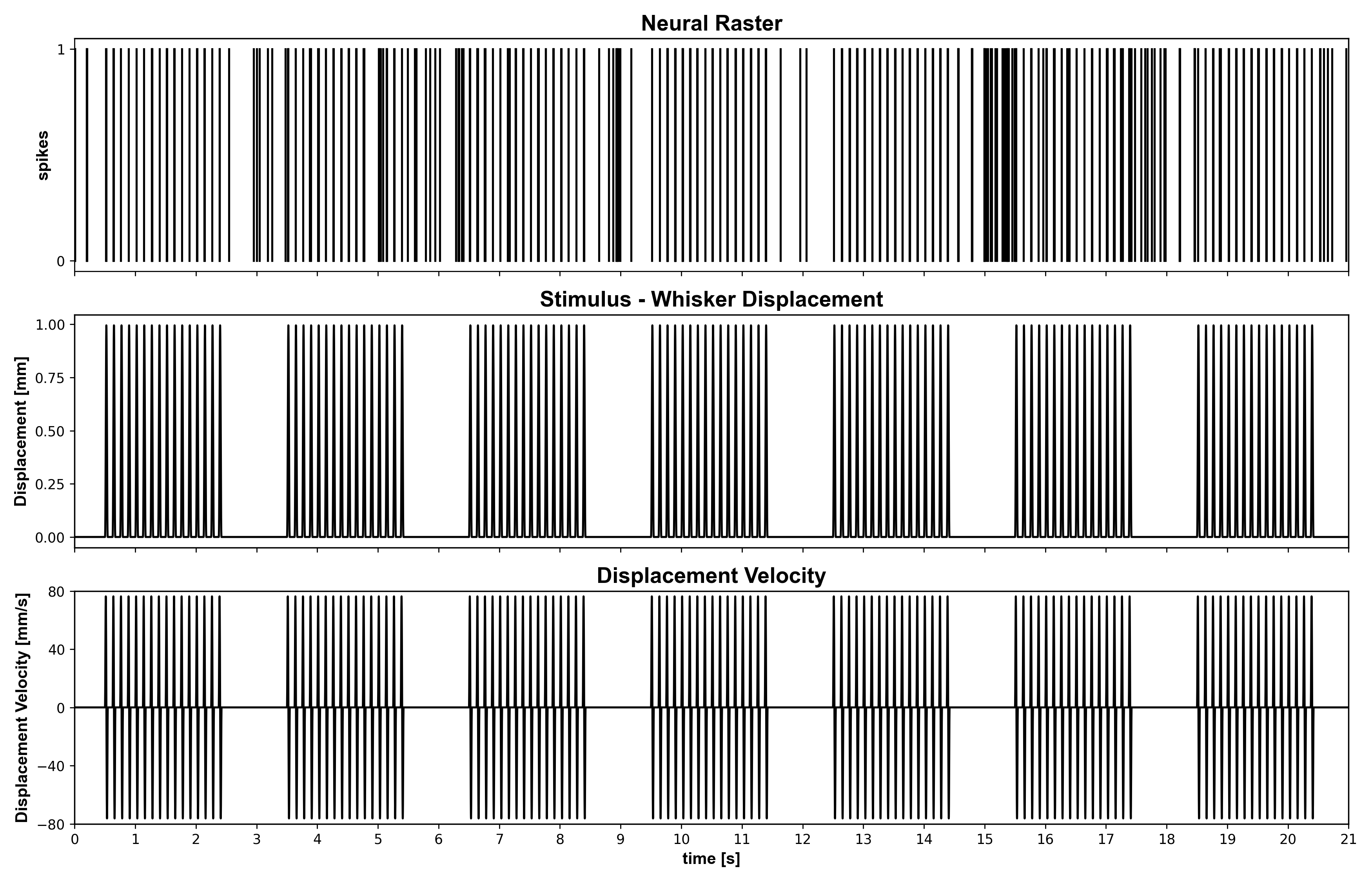
Expected figure files:
docs/figures/example02/fig01_data_overview.pngdocs/figures/example02/fig02_lag_and_model_comparison.png
Example 03: PSTH and SSGLM Dynamics Example
Question: How do PSTH and SSGLM capture within-trial and across-trial dynamics?
Run command: python examples/paper/example03_psth_and_ssglm.py
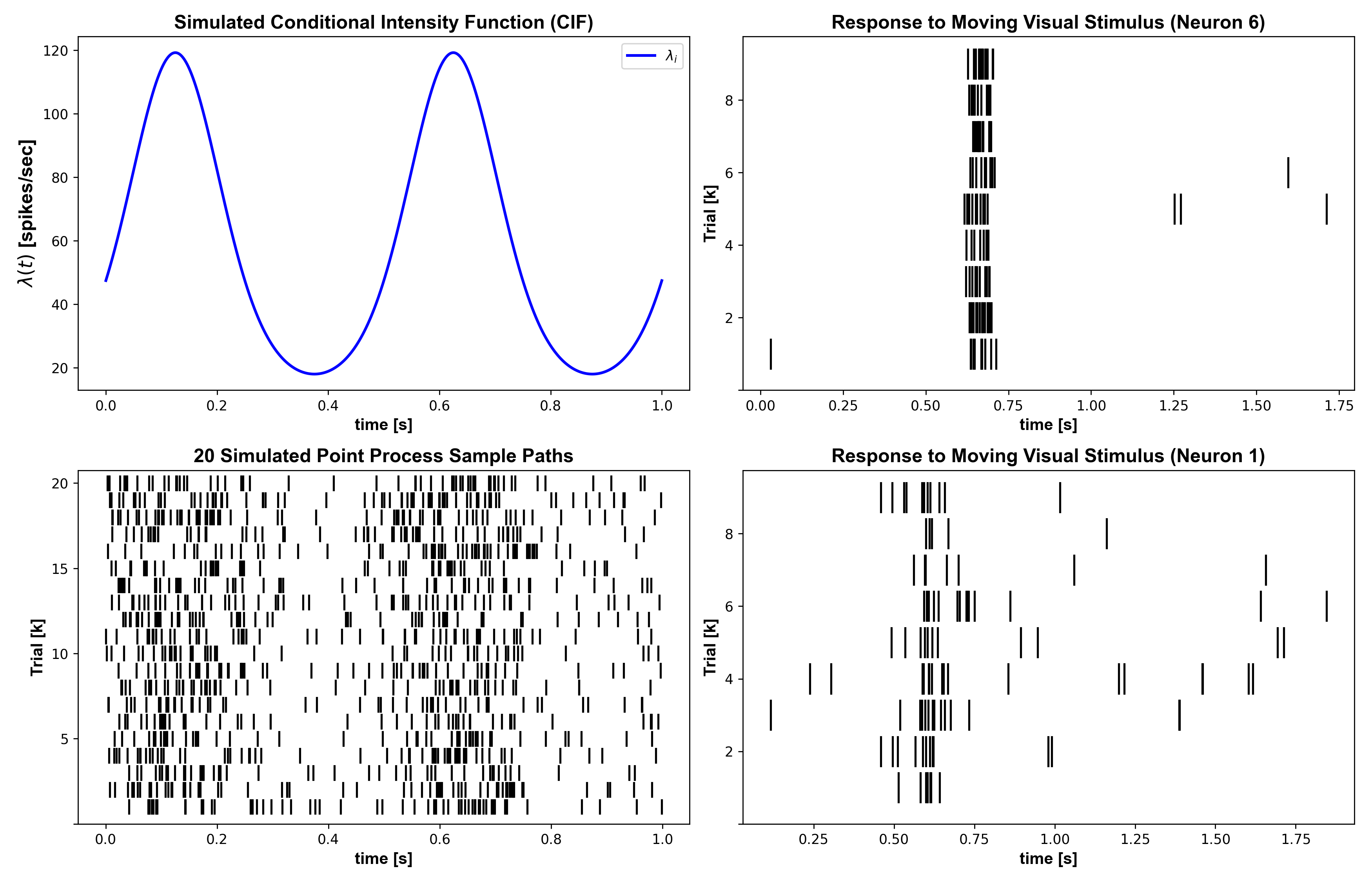
Expected figure files:
docs/figures/example03/fig01_simulated_and_real_rasters.pngdocs/figures/example03/fig02_psth_comparison.pngdocs/figures/example03/fig03_ssglm_simulation_summary.pngdocs/figures/example03/fig04_ssglm_fit_diagnostics.pngdocs/figures/example03/fig05_stimulus_effect_surfaces.pngdocs/figures/example03/fig06_learning_trial_comparison.png
Example 04: Place-Cell Receptive Fields (Gaussian vs Zernike)
Question: Which receptive-field basis (Gaussian vs Zernike) better fits place cells?
Run command: python examples/paper/example04_place_cells_continuous_stimulus.py
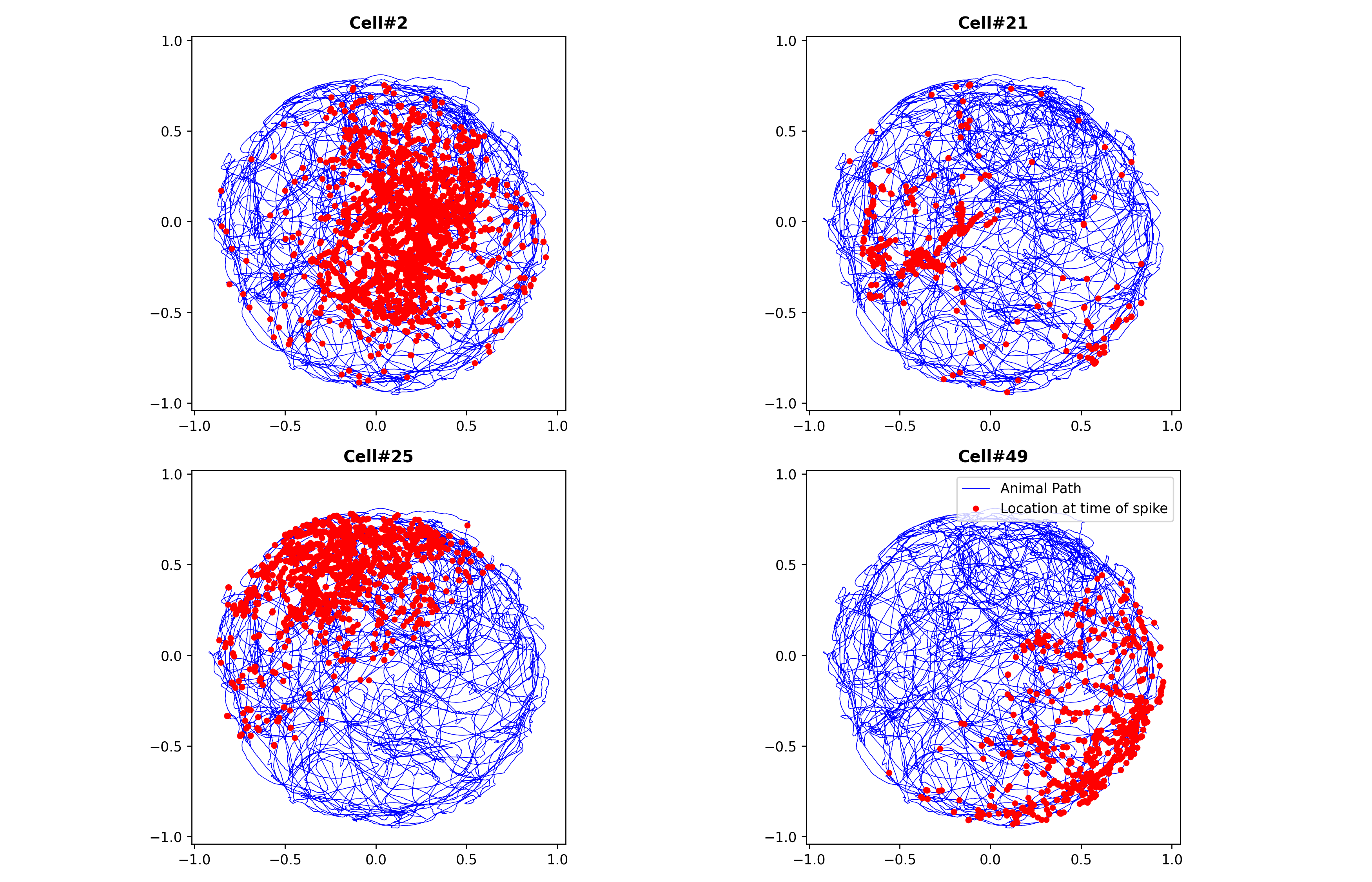
Expected figure files:
docs/figures/example04/fig01_example_cells_path_overlay.pngdocs/figures/example04/fig02_model_summary_statistics.pngdocs/figures/example04/fig03_gaussian_place_fields_animal1.pngdocs/figures/example04/fig04_zernike_place_fields_animal1.pngdocs/figures/example04/fig05_gaussian_place_fields_animal2.pngdocs/figures/example04/fig06_zernike_place_fields_animal2.pngdocs/figures/example04/fig07_example_cell_mesh_comparison.png
Example 05: Stimulus Decoding With PPAF and PPHF
Question: How well do adaptive/hybrid point-process filters decode stimulus and reach state?
Run command: python examples/paper/example05_decoding_ppaf_pphf.py
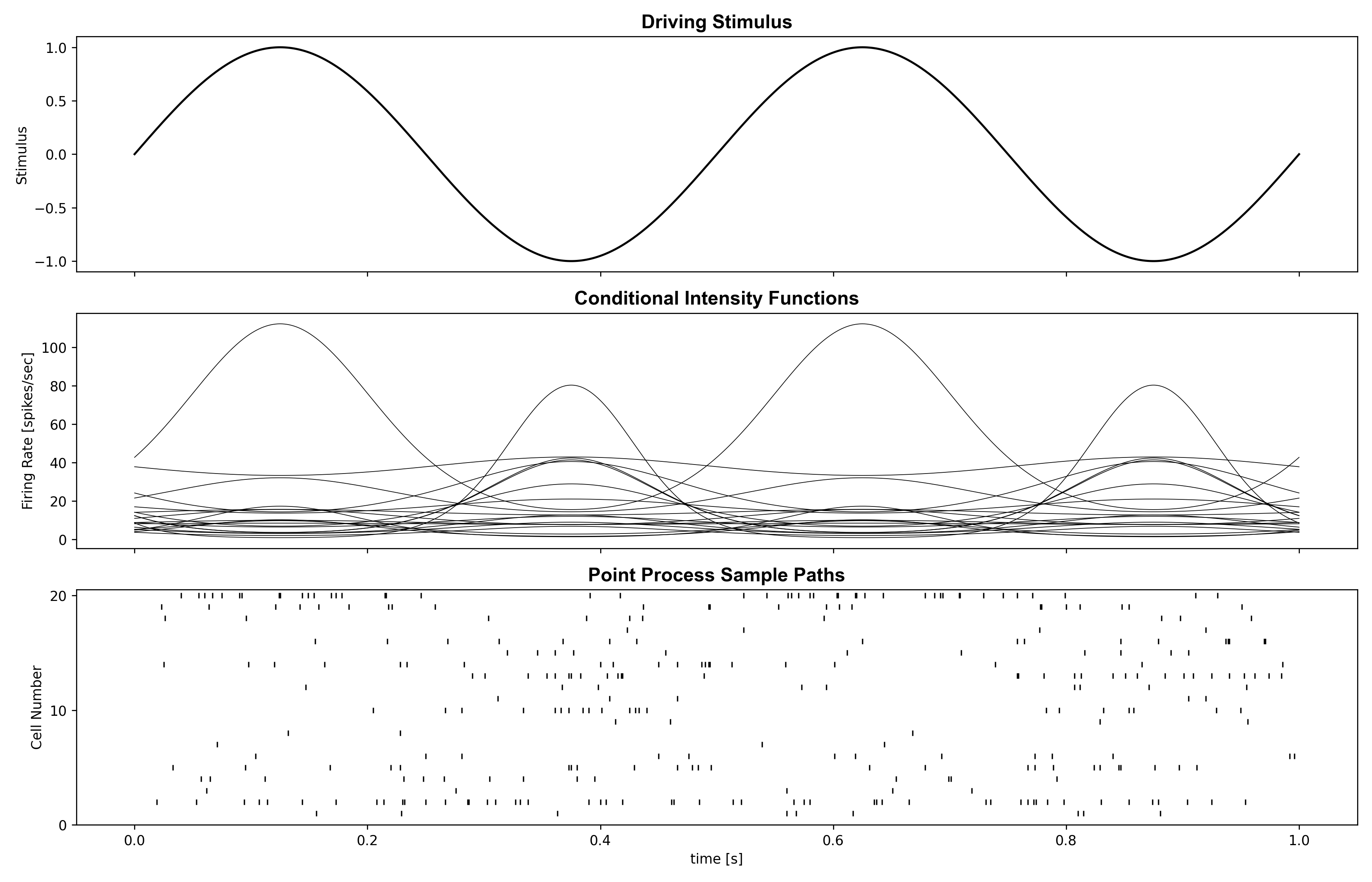
Expected figure files:
docs/figures/example05/fig01_univariate_setup.pngdocs/figures/example05/fig02_univariate_decoding.pngdocs/figures/example05/fig03_reach_and_population_setup.pngdocs/figures/example05/fig04_ppaf_goal_vs_free.pngdocs/figures/example05/fig05_hybrid_setup.pngdocs/figures/example05/fig06_hybrid_decoding_summary.png